Theoretical analysis of adaptive evolution
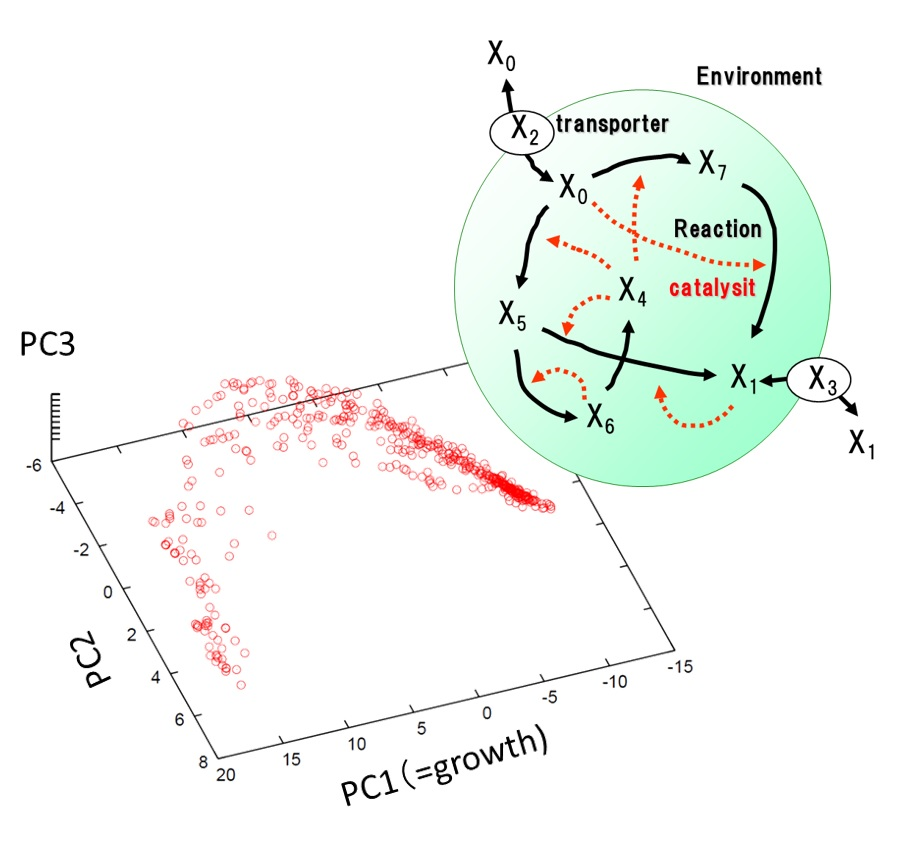
Biological systems have a high dimensional of dynamics that consists of diverse components. How can these complex systems evolve stably against environmental changes? Using theoretical analysis and computer simulations of cell models, we explored the universal characteristics of plasticity and robustness in the evolutionary processes. As a result, we found that there is a positive correlation between phenotypic fluctuation and evolutionary response independent of genome mutation. Furthermore, we found that the cell state is constrained on low-dimensional dynamics to satisfy the evolutionary stability, which was verified by experimental data. These results suggest that even under random genetic changes, there is a deterministic direction in the evolution of phenotypes. We are proceeding to analyze such low-dimensional dynamics both theoretically and experimentally.
Formation of dominant mode by evolution in biological systems, Phys. Rev. E (2018)
High-throughput bacterial experimental evolution
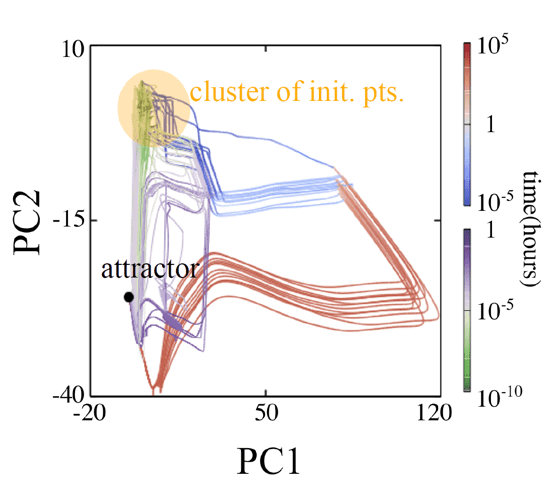
To understand the evolvability of biological systems, we performed the experimental evolution of E. coli under various environments. Using a fully automated system designed for experimental evolution (right figure), we have completed evolution experiments under 95 different stress substances, including inhibitors of cell wall synthesis and protein synthesis, as well as acids, alkalis, and heavy metals. The stress-tolerant strains obtained from these experiments were subjected to transcriptome analysis for gene expression profiles and genome resequencing analysis. The analysis of these data using appropriate machine learning methods showed that the phenotypic evolution of E. coli is constrained by relatively low-dimensional dynamics. This is in good agreement with the results of the theoretical studies mentioned above. We are now analyzing the nature of the evolvability in detail by applying dynamic feedback control to the selection environment.

Theoretical analysis of metabolic dynamics
The metabolic system consists of a network of chemical reactions by which organisms produce their components and energy for self-maintenance based on nutritional substances taken in from the environment. Most of the theoretical research on metabolism has been based on the steady-state assumption, i.e., that the concentration of chemical substances inside the cell does not change over time. In an environment where nutrients are abundantly supplied, and microbial cells grow rapidly, theories based on this assumption have been able to predict experimental results with some accuracy. In recent years, however, there have been reports of phenomena that cannot be explained by the steady-state assumption, such as the fact that the behavior of cells in a starvation environment without nutrients depends on the past culture environment. To understand the behavior of microbial cells in such environments, we are modeling the metabolic reaction system of E. coli using a set of differential equations and comprehensively investigating how the cells behave under various environments by using dynamical systems and network theory.
Experimental analysis of evolvability of gene expression
What determines the rate of gene expression evolution? Recent studies have theoretically predicted that the rate of expression evolution varies from gene to gene and is positively correlated with fluctuation in gene expression (stochastic variation without environment or mutation) and environmental responsiveness. This theoretical prediction has attracted much attention in recent years because of its implications for evolution's iterative, directional, and predictive nature. Still, rigorous experiments have not yet demonstrated it. Therefore, we are attempting to test the theoretical predictions by quantifying the expression levels of individual genes in E. coli in terms of fluctuations, environmental responsiveness, response to mutation, and rate of evolution. So far, we have succeeded in demonstrating a positive correlation between fluctuations and environmental responsiveness, and we are now quantifying the rate of evolution.
Reconstruction of genome evolution in symbiotic bacteria
In the process of evolution from free-living to parasitic bacteria and then to an intracellular symbiont, a transient explosion in the number of transposons in the genome is known to occur. However, it is not well understood how this explosive increase affects the evolution of symbiosis. This is because it is very difficult to isolate and analyze many natural symbiotic bacteria, which are too dependent on their symbiotic partners. Therefore, we are attempting to reconstruct the process of genome evolution in symbiotic bacteria using E. coli, which is easy to cultivate and analyze in the laboratory. So far, we have succeeded in experimental evolution that dramatically increases the number of highly active transposons, and we are currently analyzing this process.
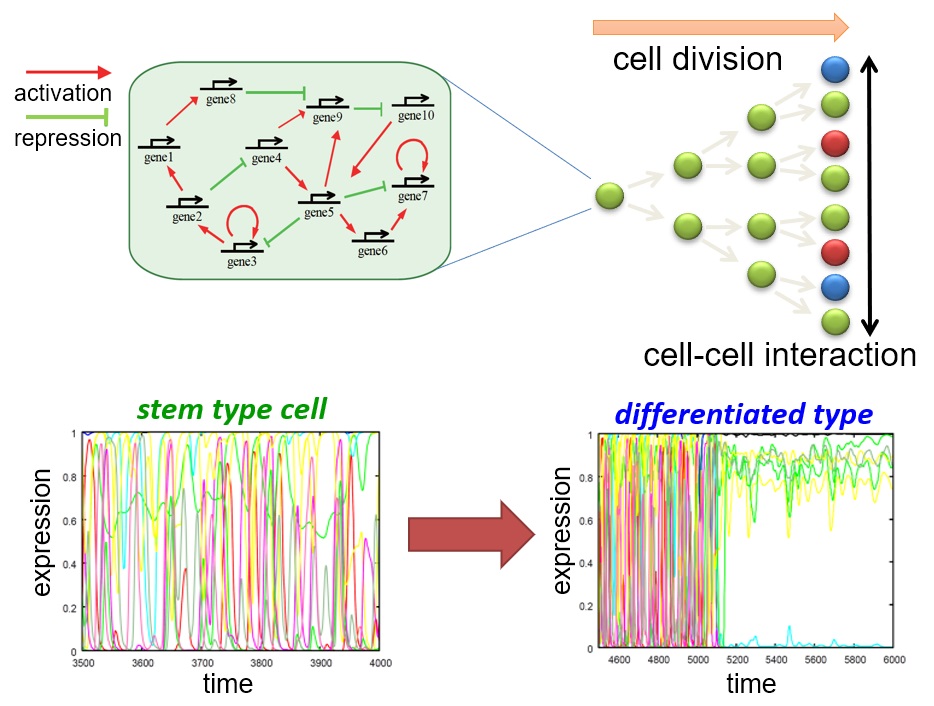
Robustness, hierarchy, and irreversibility in cell differentiation
The developmental process of multicellular organisms generally involves the differentiation to various cell types from a small number of undifferentiated stem cells. What are the mechanisms underlying this irreversible and hierarchical differentiation process? And how are these processes regulated to achieve robustness at the individual cell and population levels? To answer these questions, we used computer simulations to analyze the diversity and stability of states during development. As a result, we showed that the simple assumption that cells with nonlinear internal dynamics interact with each other is sufficient to explain differentiation into various states and stability of the developmental process. In particular, we proposed a prediction that differentiated stem cells have complex oscillating gene expression dynamics with a higher degree of freedom than terminal cells that have lost their differentiation potential. This prediction has been experimentally confirmed. By further developing these theoretical studies, we attempt to understand how hierarchical cell state transitions and stability at the cell population level are achieved during development.
A Dynamical-Systems View of Stem Cell Biology, Science (2012)